Kernel density estimates with pattern overlays
Source:R/geom_density_pattern.R
geom_density_pattern.RdA drop-in replacement for ggplot2::geom_density() that adds a pattern
aesthetic. The pattern is clipped to the filled area under the density
curve using the same device-independent in-R geometry as
geom_ribbon_pattern().
Usage
geom_density_pattern(
mapping = NULL,
data = NULL,
stat = "density",
position = "identity",
...,
na.rm = FALSE,
orientation = NA,
show.legend = NA,
inherit.aes = TRUE
)Arguments
- mapping
Aesthetic mappings created by
ggplot2::aes().- data
Data frame.
- stat
Statistical transformation. Default
"density".- position
Position adjustment. Default
"identity".- ...
Additional arguments passed to
ggplot2::stat_density()(e.g.bw,adjust,kernel,n).- na.rm
If
FALSE(default), missing values are removed with a warning.- orientation
Orientation of the layer. Default
NA(automatic).- show.legend
Logical. Should this layer be included in the legend?
- inherit.aes
If
FALSE, overrides the default aesthetics.
Pattern aesthetics
In addition to all aesthetics accepted by ggplot2::geom_density(), this
geom accepts:
patternCharacter name of the pattern. One of
"none","hatch","crosshatch","horizontal","vertical","dots","weave", or a custom pattern registered withregister_pattern(). Each base pattern (except"none") also has_denseand_sparsevariants (e.g."hatch_dense","dots_sparse") for pre-set tighter or looser spacing.pattern_colourColour of pattern lines/dots. Default
"black".pattern_linewidthLine width for line-based patterns. Default
1.pattern_spacingSpacing between pattern elements in millimetres. Default
5. Smaller values produce denser patterns; larger values produce sparser patterns.pattern_angleAngle in degrees for hatch patterns. Default
45.pattern_sizeDot radius in millimetres for the
"dots"pattern. Default0.5.
Limitations
orientation = "y" (horizontal densities) is not supported for the pattern
overlay. The base density area renders correctly, but the pattern is
silently skipped. Use coord_flip() on a vertical density as a workaround.
position = "stack" is supported: stacked densities are filled from 0 to
the cumulative density, and the pattern follows the same filled region.
Examples
library(ggplot2)
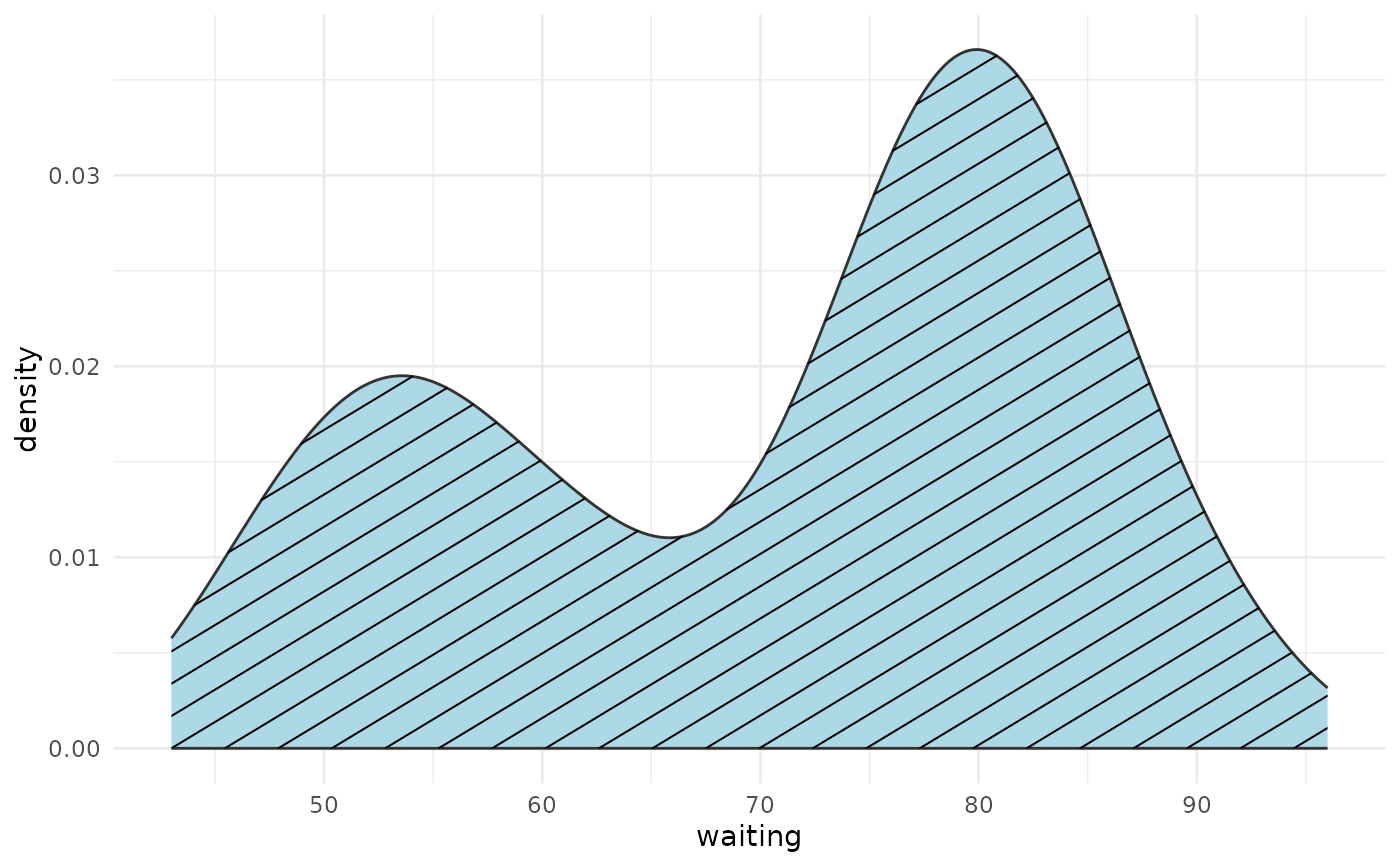
# Single density with hatch pattern
ggplot(faithful, aes(waiting)) +
geom_density_pattern(pattern = "hatch", fill = "lightblue") +
theme_minimal()
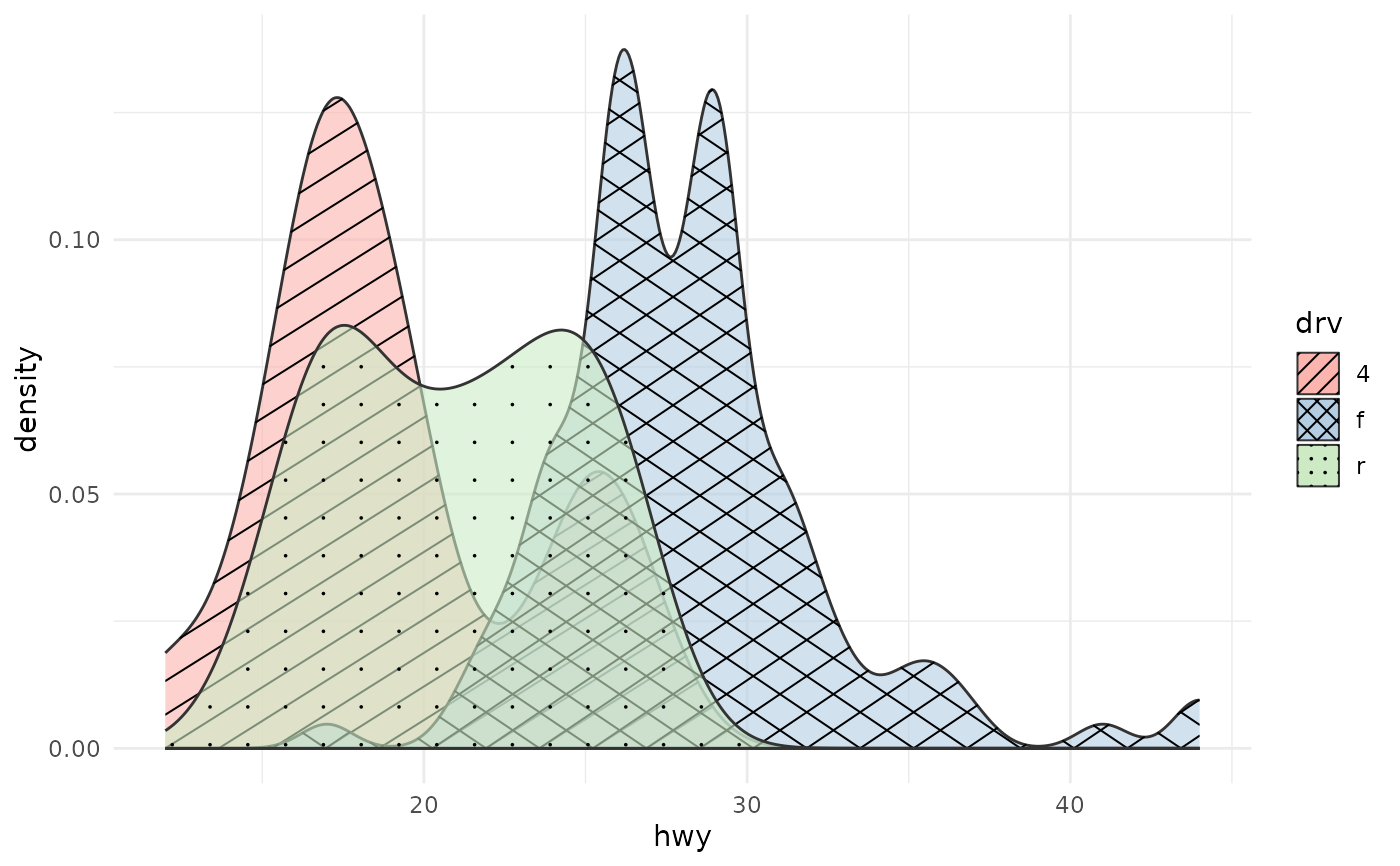
 # Multiple groups with different patterns
ggplot(mpg, aes(hwy, fill = drv, pattern = drv)) +
geom_density_pattern(alpha = 0.6) +
scale_pattern_manual(values = c("4" = "hatch", "f" = "crosshatch",
"r" = "dots")) +
scale_fill_brewer(palette = "Pastel1") +
theme_minimal()
# Multiple groups with different patterns
ggplot(mpg, aes(hwy, fill = drv, pattern = drv)) +
geom_density_pattern(alpha = 0.6) +
scale_pattern_manual(values = c("4" = "hatch", "f" = "crosshatch",
"r" = "dots")) +
scale_fill_brewer(palette = "Pastel1") +
theme_minimal()